|
In this vendor presentation given at WCBP, we introduce a new glycan fluorescent label, RapiFluor-MS, which is used to label N-linked glycans. This innovative label improves FLR and mass spectrometry signals for glycan characterization and profiling analysis. Plus - our GlycoWorks RapiFluor-MS N-Glycan Kit now allows you to finish glycan deglycosylation, labeling and cleanup in 3 steps and just 30 minutes.Learn more about this novel technology and its applications for glycosylated proteins:more of Waters' solutions for biopharmaceutical laboratories:a full video of this talk, recorded Jan. 27, 2015 at WCBP:http://www.waters.com/waters/library.htm?cid=10116552&lid=134833758. No notes for slide. Based on the observed chromatographic peak areas, response factors for fluorescence and MS detection were determined. These results indicate that RapiFluor-MS labeled glycans produce 2x higher fluorescence signal and, more astoundingly, nearly 800x greater MS signal than N-glycans labeled with Instant AB.
In this slide, deconvoluted mass spectra are presented for a mAb subjected to various conditions. The top spectrum shows the mAb before any treatment. The middle spectrum after treatment with only deglycosylation conditions. Lastly, the bottom spectrum after our combined approach of surfactant-based heat denaturation and Rapid PNGase F deglycosyaltion.
Chlorogenic acids (CGA) have been profiled in the leaves of Momordica balsamina, Momordica charantia, and Momordica foetida. All three species were found to contain the trans and cis isomers of 4-acyl para-coumaroylquinic acid ( pCoQA), caffeoylquinic acid (CQA), and feruloylquinic acid (FQA). To the best of our knowledge, this is the first report of pCoQA and FQA and their cis isomers in these Momordica species.
These profiles were obtained by a newly developed UPLC-qTOF-MS method based on the in-source collision induced dissociation (ISCID) method optimized to mimic the MS 2 and MS 3 fragmentation of an ion trap-based MS. The presence of the cis isomers is believed to be due to high UV exposure of these plants. Furthermore, the absence of the 3-acyl and 5-acyl CGA molecules points to a metabolic mark that is unusual and represents a very interesting biochemical phenotype of these species. Our optimized ISCID method was also shown to be able to distinguish between the geometrical isomers of all three forms of CGA, a phenomenon previously deemed impossible with other common mass spectrometry systems used for CGA analyses. IntroductionChlorogenic acids (CGA) are natural antioxidants and one of the most abundant polyphenols in the human diet. These molecules have multiple functions in plants and in most cases; they are produced as a part of the defense mechanism response against environmental stresses triggered by microbial pathogens, mechanical wounding, and direct exposure to UV or visible light ,. To avoid any ambiguity, in the current study, CGA refer to any molecule which is formed from an ester-bonding between one or more cinnamic acids ( p-coumaric, caffeic, and ferulic acid) and quinic acid, giving thus rise to p-coumaroylquinic acid ( pCoQA), caffeoylquinic acid (CQA), and feruloylquinic acid (FQA), respectively ,.
The metabolic profiles generated by LC/MS allowed the reconstruction of the flavonoid biosynthetic pathways, which revealed a constitutive character for herbivory resistance in the resistant genotype IAC-17 and a metabolic regulation for the rechanneling of Quercetin, Kaempferol and Genistein conjugates in soybean.
In nature, the trans isomers are currently known to be more abundant than the cis isomers; and the latter are known to accumulate in tissues/cells subjected to intense UV irradiation.From analytical point of view, specialized platforms and approaches are indeed needed to confidently and accurately annotate these molecules, if present in a particular material (e.g., plant). Clifford et al. proposed a mass spectrometry (MS) based method for identification of CGA, underlining the hierarchical fragmentation of these molecules in an MS experiment. Such observed fragmentation patterns pointed to differential behaviors of the CGA isomers during an MS analysis. Even though it has been shown that MS is capable of differentiating the regional isomers, formed from differential positional acylation, this platform is reportedly unable to distinguish between the geometrical isomers ( cis and trans), with the latter resulting in very similar MS-fragmentation patterns. The only way to putatively characterise these geometrical isomers so far is by their differential retention time during a chromatographic separation. In a reverse chromatography, the cis-3-acyl and cis-4-acyl molecules elute before their trans counterparts whilst the opposite is the case for 5-acyl molecule.It is indeed clear that the CGA molecules offer a very interesting analytical challenge that still needs more investigation, chromatographically and at the MS level, for more distinct annotation of different molecular forms of these compounds.
In the scientific community, various researchers are therefore spending most of their time designing reliable methods to characterize these molecules (and their associated derivatives) from different sources of plant materials. In the current study, a newly developed UPLC-Q-TOF-MS method, which is based on the in-source collision induced dissociation, is evaluated for the characterization of chlorogenic acid content of the three Momordica species growing in the northern part of South Africa.Momordica genus (Cucurbitaceae) is known to have high nutritious values and ethnopharmacological activities.
Plants from this genus are known to possess a very bitter taste which has been attributed to the cucurbitacins. These compounds have been shown to exhibit several potent biological activities, namely, cytotoxic, hepatoprotective, anti-inflammatory, cardiovascular, antidiabetic, and antiparasitic –. To the best of our knowledge, there is very little work done on the other classes of compounds (with potential biological activities) that are possibly present in Momordica. In South Africa, Momordica species are used as personal prescription medicine to control sugar diabetes.
Several studies have indicated that the spectrum of naturally occurring compounds, with biological activities, includes various classes such as phenolics, carotenoids, and flavonoids. CGA have been previously shown to possess antidiabetic properties. Thus, the current study is exploring the presence of chlorogenic acids in Momordica found in the northern part of South Africa, with a focus on the Q-TOF-MS-based annotation of the CGA molecules, explaining their fragmentation patterns as a tool to discriminate different forms of CGA. Thus, to achieve the CGA characterization, the in-source collision induced dissociation aided mass spectrometry was used. Partial optimization of the ISCID method for the discrimination of geometrical isomers of CGA is also presented. Metabolites ExtractionThe leaves of the three Momordica species were air-dried using an oven operating at 37°C for three consecutive days.
The dried leaves were crushed using a pestle and mortar to a fine to coarse powder with a relative (homogeneous) size. Metabolites were extracted from the crushed leaves (2 g) using 80% MeOH (20 mL). After shaking at room temperature (25°C) for 30 min, the tissue debris was removed by means of centrifugation at 5000 ×g for 10 min. The supernatant was transferred to a new clean tube and dried to at least 1.0 mL using the rotary vapor apparatus at 55°C under vacuum. The 1 mL extract was subsequently dried to completeness using the speed vacuum concentrator centrifuge at 55°C.
The resulting pellet was reconstituted in 1.0 mL of 50% aqueous MeOH and filtered through 0.22 µm nylon filters. The filtered 1.0 mL extract was then analysed on an UPLC-MS system. UPLC-MS AnalysesChromatographic separation (of the 1 mL extract) was performed on an ACQUITY UPLC system (Waters Corporations, Milford, MA) using a conditioned autosampler at 4°C. One (1) µL of the extracts was separated on a Waters BEH C8 column (150 mm × 2.1 mm, 1.7 µm), thermostatted at 60°C. A binary solvent mixture was used consisting of water (Eluent A) containing 10 mM formic acid (natural pH of 2.3) and acetonitrile (Eluent B) containing 10 mM formic acid.
The initial conditions were 98% A at a flow rate of 0.4 mL/min and were maintained for 1 minute, followed by multiple gradients to 5% A at 26 minutes. The conditions were kept constant for 1 minute and then changed to the initial conditions. The total chromatographic run time was 30 minutes.
High definition mass spectrometry was performed on a Waters SYNAPT G1 Q-TOF system in V-optics and operated in electrospray negative mode to enable detection of phenolic compounds. Leucine enkephalin (50 pg/mL) was used as reference lock mass calibrant to obtain typical mass accuracies between 1 and 5 mDa. The optimal conditions of analysis were as follows: capillary voltage of 2.5 kV, the sampling cone at 30 V, and the extraction cone at 4 V. The scan time was 0.1 seconds covering the 100 to 1000 Dalton mass range. The source temperature was 120°C and the desolvation temperature was set at 450°C. Nitrogen was used as the nebulisation gas at a flow rate of 700 L/h.For targeted approach, the CGA were profiled by collecting MS/MS (typical MS 2) data of the masses of interest, for CQA at m/z 353, pCoQA at m/z 337, and FQA at m/z 367. To achieve the diagnostic fragmentation patterns reported by Clifford et al.
of these compounds, the trap collision energy (3–60 eV) and the cone voltage (10–100 V) were experimentally optimised until a stable fragmentation pattern was achieved characterized by the formation of the following stable ions: Q 1Quinic acid-H − at m/z 191, C 1Caffeic acid-H − at m/z 179, Q 2Quinic acid-H-H 2O − at m/z 173, and C 2Caffeic acid-H-CO 2 − at m/z 135. The exact masses, m/z, (and their respective elemental formulae) for some of these ions detected in the current study are indicated in.
Most importantly, the ISCID method was optimized to result in a more stable m/z 191 (Q 1) in order to generate this ion upfront inside the source because it was noted, from our preliminary results, that it was ubiquitous in all regional and geometrical isomers. All the acquisition and analysis of data were controlled by Waters MassLynx v4.1 software (SCN 704).
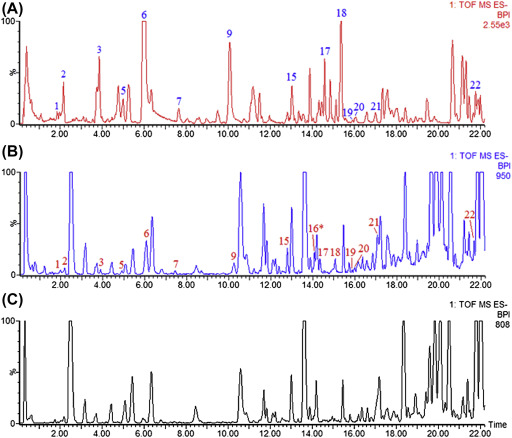
Results and DiscussionBefore embarking on the targeted analysis for profiling chlorogenic acids in the above-mentioned three Momordica species (the focus of this study), the UPLC-MS-based untargeted analyses of methanol extracts of the three species revealed inherent differential metabolite content as indicated by principal component analysis (PCA) models of the data (results not shown). This different clustering in PCA models points, thus, to an imminent need for more investigation in the metabolite content of these three species. This untargeted analyses indicated putatively the presence of chlorogenic acids in the methanol extracts of the three Momordica species under study.Though chlorogenic acid molecules are naturally found in most plants, their accurate identification is often very challenging analytically due to the chemical structural forms (in terms of regional and geometrical isomerism) of these compounds. Clifford et al.
have developed a more reliable and validated ion trap (IT)-MS-based method for annotation of CGA molecules, exploring hierarchical fragmentation of a molecule in multistage mass spectrometry (MS n) analyses. The latter are currently possible only on an ion trap MS system. However, with the increased usage of Q-TOF instruments in plant metabolomics (and natural products) research –, it is imperatively important to develop and optimize a Q-TOF based method (for the identification of CGA molecules from plant extracts) which gives accurate results similar to those achieved by an IT-MS platform.Thus, to illustrate in more detail our Q-TOF based method for the annotation of CGA molecules, it suffices to highlight some aspects regarding the consistently observed fragmentation patterns of these compounds in MS analyses (MS tuned as detailed in ). For instance, in regard to the annotation of CQA molecules, the 4-acylated CQA was identified by the presence of a base peak at m/z 173 (Q 2) (, ), which is in agreement with the results generated from an ion trap-MS-based method. The analyses indicated that the 3- and 5-CQA molecules are not present in the Momordica species studied. Normally, the MS fragmentation patterns of the 3- and 5-CQA molecules show m/z 191 as the base peak for these molecules , which is different from the 4-acylated CQA that has the base peak at m/z 173 (identified in this study:, ).
Furthermore, the 3-CQA is distinguished from the 5-CGA by the presence of a base peak at m/z 191 with another secondary peak at m/z 179 whilst the 5-CQA only shows base peak at m/z 191 without a secondary peak at m/z 179. Using a similar approach, other CGA molecules that were confidently annotated in this study are 4- pCoQA and 4-FQA molecules. All these observations agree with the ion trap MS results.
The structures of chlorogenic acids detected in the Momordica species.All the annotated CGA molecules in this study, that is, 4-CQA, 4- pCoQA, and 4-FQA, show two ion peaks in their chromatographic separation , which is an indication for the presence of geometrical isomers ( trans and cis). To date, the ion trap-MS-based method has shown that the fragmentation patterns of both cis and trans chlorogenic acid molecules are very similar, hence the two could not be distinguished by MS analyses.
However, in this study, the MS/MS data of pCoQA, CQA, and FQA, from our ISCID based method, revealed some clear differences between the trans and cis isomers of 4-CQA molecule. Here, both the cis and trans isomers have the base peak at m/z 173 quinic acid-H-H 2O (C 1) and a secondary fragment at m/z 191 (Q 1); however, in all cases, the intensity of the m/z 191 fragment (Q 1) is higher in the trans than in the cis isomer. It can be postulated that this might be due to energy distribution differences, on quantum level, which is causing these molecules/isomers to behave differently under similar MS conditions. This could be MS-fragmentation signatures that would aid in distinguishing these two geometrical isomers of the 4-CQA molecule.
Representative mass spectra showing the major fragmentation differences at m/z 191 noted between trans and cis isomers of all CGA detected in Momordica species. The exact mass of m/z 191 and its proposed elemental formula are indicated in.Our on-going research, as seen from this study , shows that Q-TOF-MS has an extra ability of differentiating geometrical isomers. Furthermore, we note that the obtained fragmentation patterns of CGA molecules from the Momordica species (Figures and ) are in agreement with those of already published data generated from ion trap MS systems (with no MS-based differentiation of cis and trans isomers) and were confirmed using MS fragmentation of CGA from alcoholic cider and coffee extracts (considered as CGA standards for their known CGA content) which were analysed contemporaneously.As previously stated, in an attempt to distinguish the geometrical isomers of CGA, Clifford et al. concluded that ion trap MS shows similar fragmentation patterns between trans and cis isomers. To overcome this inability of IT based MS, Clifford et al.
opted for chromatographic retention parameters of these molecules, observing that the cis isomers are capable of forming intramolecular hydrogen bonding which is minimal/absent in trans isomers. Based on their finding, it was postulated that, in reversed phase chromatography, the cis isomers for 3-acyl and 4-acyl elute before their trans counterparts and the opposite was observed in the case for 5-acyls. To date, this is the basis in which CGA geometrical isomers are annotated: difference in elution times in reversed phase chromatography.Here, we suggest thus an MS-based method that differentiates CGA geometrical isomers ( cis and trans).
Similar experiments were conducted in three more CGA sources, namely, the Moringa oleifera leaves methanolic extracts, Nicotiana tabacum cell cultures methanolic extracts, and alcoholic ciders, resulting substantially in the same observation: differentiation of cis and trans CGA molecules based on their behavior in the MS. ConclusionMomordica species, investigated in this study, contain a range of chlorogenic acids, particularly the 4-caffeoylquinic acid, 4- p-coumaroylquinic acid, and 4-feruroylquinic acid.
However, the absence of other regional isomers (3- and 5-acyls) was unexpected, as they are the most frequently existing forms in many plants. The presence of chlorogenic acids in Momordica species, as demonstrated in this study, warrants further investigation by looking at their roles as possible antidiabetic compounds functioning synergistically with the previously identified cucurbitacins. Thus, this study clearly shows the ability of Q-TOF-MS as a technique for metabolite annotation in a mixture of compounds, with a unique promise to distinguish between geometrical isomers (particularly in the case of CGA molecules, as indicated in this study).
Thus, fine-tuning the MS settings will definitely result in clearly distinguishable fragmentation patterns between the other chlorogenic acids geometrical isomers ( cis and trans), a phenomenon which has been deemed impossible with ion trap-based MS instruments.
Comments are closed.
|
Details
AuthorWrite something about yourself. No need to be fancy, just an overview. ArchivesCategories |
- Home
- About Brooke
- Blog
- Cara Mempercepat Windows 10
- Clave Para Windows 7 Ultimate
- Tecarta Bible For Windows
- London Thumakda Lyrics
- Hosts File Windows 10
- Download Lagu Via Vallen Selow
- Bpmn Webservice Java
- Contact
- Notability 3 Dmg
- Tudo Sobre Network Toolkit
- Tower of trample cheat
- The owl house season 1 episode 1 online
- How to increase steam download speed 2016
- Low Specs Experience Crack Premium
- Pimpandhost-net sandra orlow nude
- Ev3 drag racing mod
- Jenny mod minecraft download 1-15-2
- Dex vocaloid statue
- Twitter video download chrome
- Elizabethan Era Game Ball
- How to download zoom on macbook air
- Youtube download windows 11
- Usando hack en critical ops pc
- Pcsx2 1-4-0 bios download emuparadise
- Jape ja2 save game editor
- Teamviewer download free full version
- Acronis true image 2014 serial numbers
- Allplan smartparts download
- Twitter video download shortcut android
- Roblox bones c4d download
- Free paysafecard codes list dragon awakening
- Garrys mod nude model addon
- Download warriors orochi z pc english full
- Ddtank nexus 8-3
- Download spss 22 full crack
- Kenneth hagin healing youtube
 RSS Feed
RSS Feed
